Dose Profiler matter studies page
Rando Full Simulation studies
The file chain is the following:- Simulation input: /NAS_arpg/mfischetti/SimRandoLight/notr_profDATA_rando_280_15R.inp
- modify runMode = 4
- put the correct detector positioning.
- make -C plugin&&freddevel
- Run on the file produced by AnaProfi. Should compare with. /NAS_arpg/MeMs/RandoData/rando280_tracks.txt
- Produces as output something: ProfilerWeights2D.txt (check plugin for content).
- /home_arpg/mfischetti/MeMs/MLEM/localout/BP.txt
The weights definition + machinery
Here we document how the weights are defined, where is the codeHow to run simulations and produce files to apply weights:
To Run V2: -reference cilinder, necessary if the target does not start at 0cm (as reference cilinder3 does) -$FLUKA/flutil/rfluka -e fluka_FILTER_mbcil.exe -N0 -M5 filter_mbcil_1R &
$FLUKA/flutil/rfluka -e fluka_FILTER_mbcil.exe -N0 -M5 filter_mbcil_2R &
$FLUKA/flutil/rfluka -e fluka_FILTER_mbcil.exe -N0 -M5 filter_mbcil_3R &
$FLUKA/flutil/rfluka -e fluka_FILTER_mbcil.exe -N0 -M5 filter_mbcil_4R & -mb brain+bone radius_ext = 10 cm centered in (0,0,10)
$FLUKA/flutil/rfluka -e fluka_FILTER_mbfull.exe -N0 -M5 filter_mbfull_1R &
$FLUKA/flutil/rfluka -e fluka_FILTER_mbfull.exe -N0 -M5 filter_mbfull_2R &
$FLUKA/flutil/rfluka -e fluka_FILTER_mbfull.exe -N0 -M5 filter_mbfull_3R &
$FLUKA/flutil/rfluka -e fluka_FILTER_mbfull.exe -N0 -M5 filter_mbfull_4R & cat filter_mbcil*00*dat > filter_mbcil_all_TXT.dat &
cat filter_mbfull*00*dat > filter_mbfull_all_TXT.dat & ./Txt2Root -in filter_mbcil_all_TXT.dat -out filter_mbcil_all.root & Then in ../ApplyWeights: see ListApplyWeights _mbfull.sh (the first part) ->to have reference hZ distr Then: cd ../V5 ln -s ../V2/filter_mbfull_all_TXT.dat secondary_TXT.dat and run FLUKA on new ray*inp files (same full geo, different beam (10TeV muon+) and physics card)
NB: check the source for reso!
$FLUKA/flutil/rfluka -e fluka_RAY.exe -N0 -M1 ray_filter_mbfull_reso0 &
$FLUKA/flutil/rfluka -e fluka_RAY.exe -N0 -M1 ray_filter_mbfull_reso1 &
$FLUKA/flutil/rfluka -e fluka_RAY.exe -N0 -M1 ray_filter_mbfull_reso2 &
$FLUKA/flutil/rfluka -e fluka_RAY.exe -N0 -M1 ray_filter_mbfull_reso3 & ./Txt2Root -in ray_filter_mbfull_reso0001_TXT.dat -out ray_filter_mbfull_reso0.root &
./Txt2Root -in ray_filter_mbfull_reso1001_TXT.dat -out ray_filter_mbfull_reso1.root &
./Txt2Root -in ray_filter_mbfull_reso2001_TXT.dat -out ray_filter_mbfull_reso2.root &
./Txt2Root -in ray_filter_mbfull_reso3001_TXT.dat -out ray_filter_mbfull_reso3.root & Then in ApplyWeights: see ListApplyWeights _mbfull.sh (second part) to produce the weighted Z distr - Compare the distributions with TestWeighs _mb.cc o quello che hai <3
The mental ball configurations
Beam: 12C ions, energy: 300 MeV /u, spatial distribution: gaussian with FWHM_x,y = 0.5 cm direction: positive z axis, starting point:(0,0,4) cm 1)*mbfull, REFERENCE distr: mbcil* Mental Ball: sphere with rmin = 9.6 cm, rmax = 10 cm, inner: BRAIN (ICRU), outer: BONE (ICRU) center:(0,0,15) cm - fluka region number of detectors (to be implemented in ApplyWeightsLib.h): {3,4,5,6,7,8} REFERENCE distr: mbcil = MB with same geo, material: water, cut with cilinder of r = 1.5 cm 2)*mbimp, REFERENCE distr: mbimp_cil* Mental Ball: sphere with rmin = 9.6 cm, rmax = 10 cm, inner: BRAIN (ICRU), outer: BONE (ICRU) center:(0,0,15) cm Implant: sphere, r = 1cm, material: AIR, center: (0.,5.,16.) - fluka region number of detectors (to be implemented in ApplyWeightsLib.h): {0,0,0,3,4,5} NB: the 60deg detectors are cut in order to have just a portion of ~ 20 x 20 cm2 ; no 90deg detectors in this configuration REFERENCE distr: mbimp_cil = MB with same geo, material: water, cut with cilinder of r = 1.5 cm (as in sim 1) ); detectors in same config of mbimp sim 3)*mbimp2, REFERENCE distr: mbimp_cil* Mental Ball: sphere with rmin = 9.6 cm, rmax = 10 cm, inner: BRAIN (ICRU), outer: BONE (ICRU) center:(0,0,15) cm Implant: sphere, r = 2cm, material: AIR, center: (0.,5.,16.) - fluka region number of detectors (to be implemented in ApplyWeightsLib.h): {0,0,0,3,4,5} NB: the 60deg detectors are cut in order to have just a portion of ~ 20 x 20 cm2 ; no 90deg detectors in this configuration REFERENCE distr: mbimp_cil = MB with same geo, material: water, cut with cilinder of r = 1.5 cm (as in sim 1) ); detectors in same config of mbimp sim 4)*mbimp3, REFERENCE distr: mbimp_cil* Mental Ball: sphere with rmin = 9.6 cm, rmax = 10 cm, inner: BRAIN (ICRU), outer: BONE (ICRU) center:(0,0,15) cm Implant: sphere, r = 1cm, material: ADIPOSE TISSUE, center: (0.,5.,16.) - fluka region number of detectors (to be implemented in ApplyWeightsLib.h): {0,0,0,3,4,5} NB: the 60deg detectors are cut in order to have just a portion of ~ 20 x 20 cm2 ; no 90deg detectors in this configuration REFERENCE distr: mbimp_cil = MB with same geo, material: water, cut with cilinder of r = 1.5 cm (as in sim 1) ); detectors in same config of mbimp sim Mental ball with inset on beam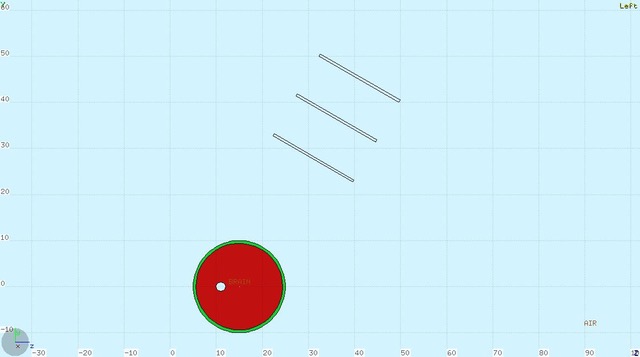
Comments
| I | Attachment | History | Action | Size | Date | Who | Comment |
|---|---|---|---|---|---|---|---|
| |
MBcil_0cm.png | r1 | manage | 111.0 K | 2018-03-01 - 16:05 | AlessioSarti | |
| |
MBcil_0cm_zoom.png | r1 | manage | 50.4 K | 2018-03-01 - 16:05 | AlessioSarti | |
| |
MBfull_0cm.png | r1 | manage | 111.0 K | 2018-03-01 - 16:05 | AlessioSarti | |
| |
MBfull_0cm_zoom.png | r1 | manage | 64.7 K | 2018-03-01 - 16:05 | AlessioSarti | |
| |
MBimpB_5cm_YZ.jpeg | r2 r1 | manage | 24.1 K | 2018-03-01 - 16:03 | AlessioSarti |
Topic revision: r7 - 2018-09-14 - AlessioSarti
Ideas, requests, problems regarding TWiki? Send feedback



